Smooth pdqr-function using random sampling and corresponding new_*() function.
form_smooth(f, n_sample = 10000, args_new = list())Arguments
| f | A pdqr-function. |
|---|---|
| n_sample | Number of elements to sample. |
| args_new |
Value
A smoothed version of f with the same class and
type.
Details
General idea of smoothing is to preserve "sampling randomness" as much as reasonably possible while creating more "smooth" probability mass or density function.
At first step, sample of size n_sample is generated from distribution
represented by f. Then, based on the sample, "continuous" d-function is
created with new_d() and arguments from args_new list. To account for
density()'s default behavior of "stretching range" by
adding small tails, support of d-function is forced to be
equal to f's support (this is done with form_resupport() and method
"reflect"). Output represents a "smooth" version of f as d-function.
Final output is computed by modifying "y" or "prob" column of f's "x_tbl" metadata to be proportional to values of "smooth" output at
corresponding points from "x" column. This way output distribution has
exactly the same "x" grid as f but "more smooth" nature.
See also
Other form functions:
form_estimate(),
form_mix(),
form_regrid(),
form_resupport(),
form_retype(),
form_tails(),
form_trans()
Examples
set.seed(101)
# Type "discrete"
bad_dis <- new_d(
data.frame(x = sort(runif(100)), prob = runif(100)),
type = "discrete"
)
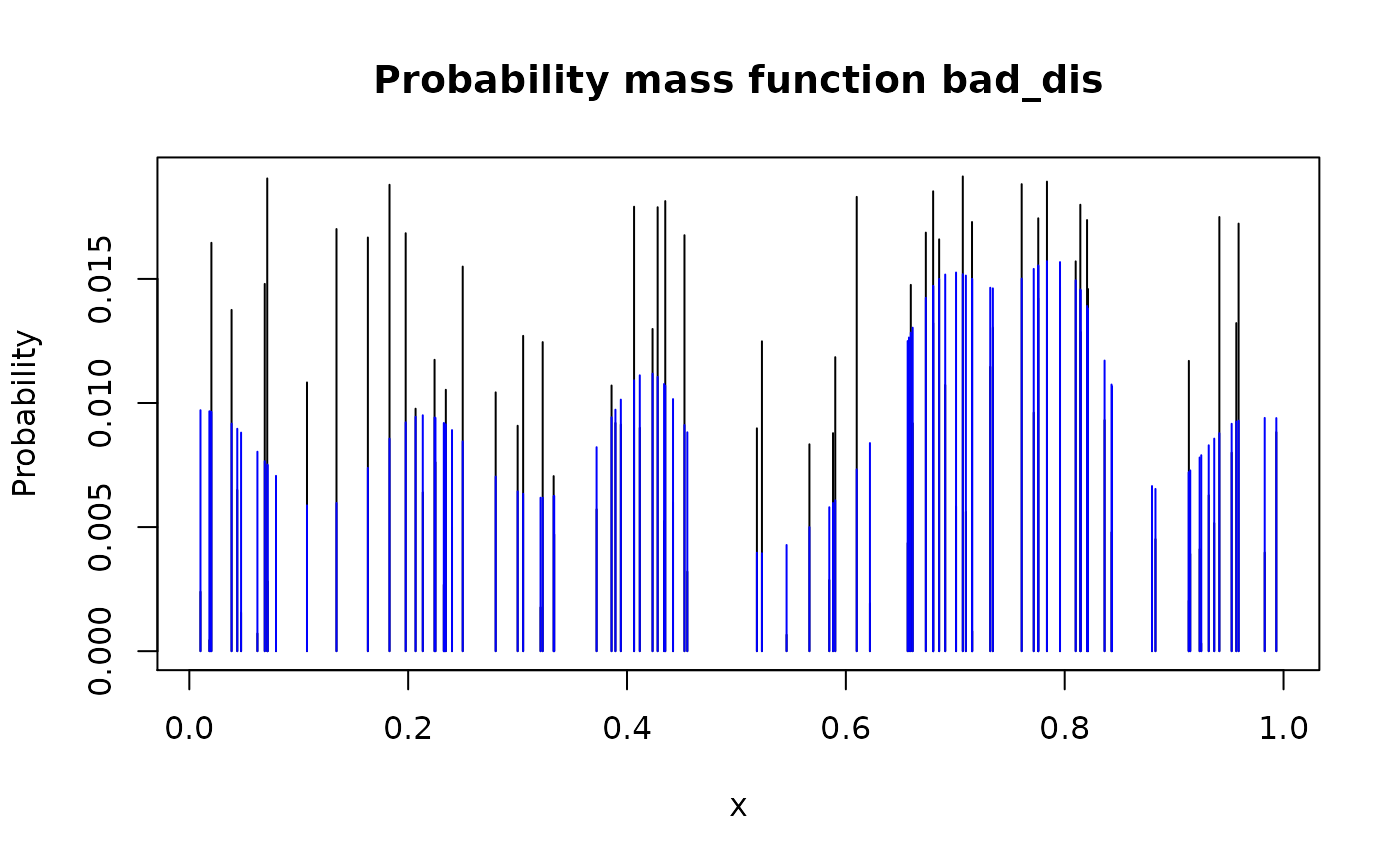
smoothed_dis <- form_smooth(bad_dis)
plot(bad_dis)
 # Type "continuous"
bad_con <- new_d(
data.frame(x = sort(runif(100)), y = runif(100)),
type = "continuous"
)
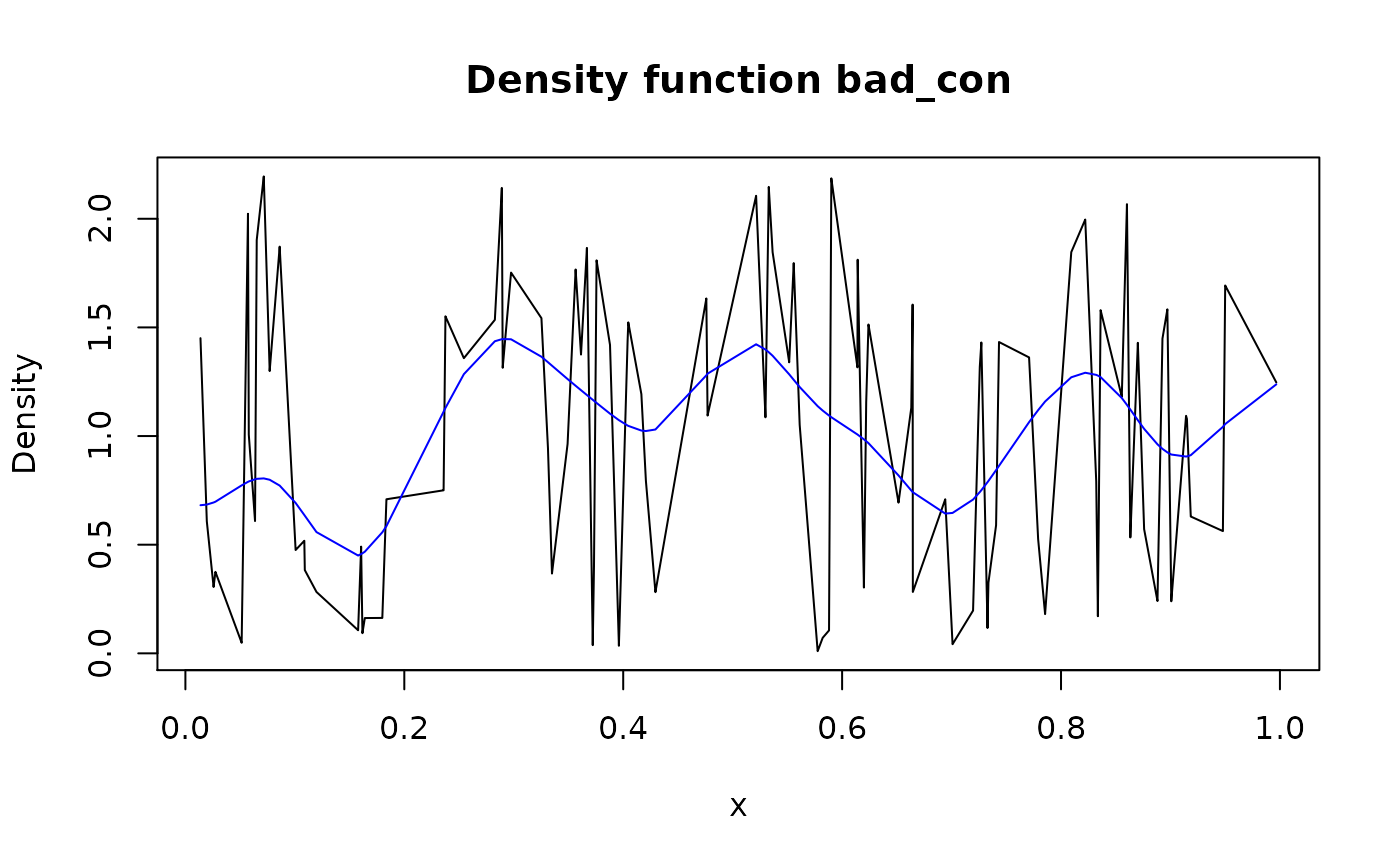
smoothed_con <- form_smooth(bad_con)
plot(bad_con)
# Type "continuous"
bad_con <- new_d(
data.frame(x = sort(runif(100)), y = runif(100)),
type = "continuous"
)
smoothed_con <- form_smooth(bad_con)
plot(bad_con)